|
|
本帖最后由 金鹏 于 2015-5-31 21:54 编辑
 N卡新包不断啊 N卡新包不断啊
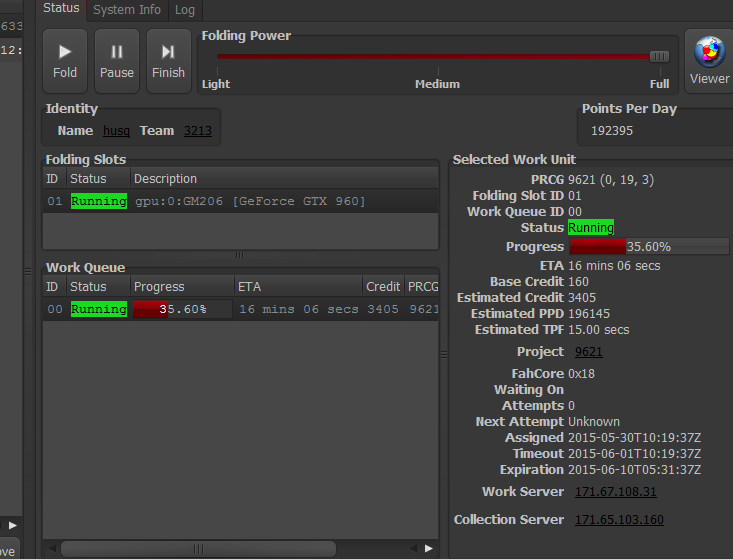
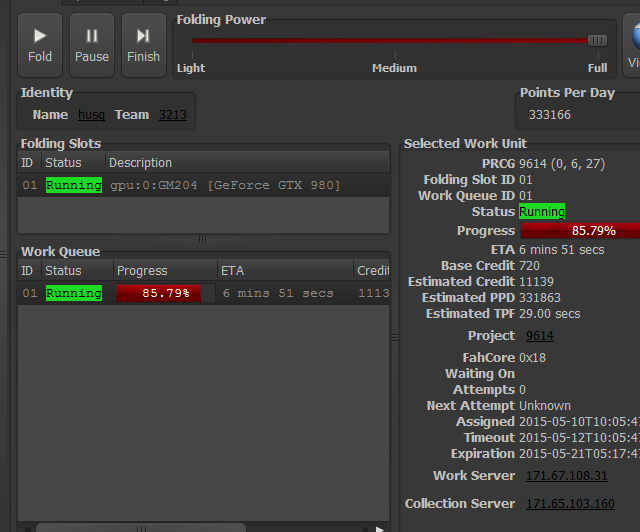
9621 and 9622 to advancedThese projects are identical in motivation to those described here: viewtopic.php?f=66&t=27666
9621 and 9622 are  mutants mutants  of 9606 and 9607, respectively. of 9606 and 9607, respectively.
SPECIFICATIONS
Uniform across projects:
x18 Nvidia GPU
8 runs
100 clones (1 us / clone)
250 gens (4 ns / gen)
10.8 deadline
2.0 timeout
0.75 k-factor
Uniquely:
project protein natoms base-credit
p9621 chignolin mutant 4128 160
p9622 chignolin mutant 5448 195 9619 and 9620 to betaThese projects are identical in motivation to those described here: viewtopic.php?f=66&t=27666
9619 and 9620 are  mutants mutants  of 9606 and 9607, respectively. of 9606 and 9607, respectively.
SPECIFICATIONS
Uniform across projects:
x18 Nvidia GPU
8 runs
100 clones (1 us / clone)
250 gens (4 ns / gen)
10.8 deadline
2.0 timeout
0.75 k-factor
Uniquely:
project protein natoms base-credit
p9619 chignolin mutant 4128 160
p9620 chignolin mutant 5448 195
Thank you  9606-9617 to FAH (NV Core 18)Dear FAH,
In molecular dynamics simulation, the system `model` refers to the equation describing the way the atoms in the simulation interact. For this project series we are running a variety of simulations using different combinations of protein and water models in order to evaluate the model-dependent differences in protein folding dynamics and resultant folded structures. Here we are focusing on 6 peptides/proteins: chignolin, trp-cage, bba, villin, ww domain, and homeodomain. Our goal is to determine which models produce the most accurate results.
SPECIFICATIONS
Uniform across projects:
x18 Nvidia GPU
8 runs
100 clones (1 us / clone)
250 gens (4 ns / gen)
10.8 deadline
2.0 timeout
0.75 k-factor
Unique for each project:
project protein natoms base-credit
p9606 chignolin 3575 130
p9607 chignolin 4720 190
p9608 trp-cage 5899 205
p9609 trp-cage 7770 310
p9610 bba 14104 690
p9611 bba 18636 1050
p9612 villin 9821 390
p9613 villin 12906 650
p9614 ww domain 15175 720
p9615 ww domain 20045 1180
p9616 homeodomain 11878 520
p9617 homeodomain 15530 830 9606-9617 to ADV (Core 18)In molecular dynamics simulation, the system `model` refers to the equation describing the way the atoms in the simulation interact. For this project series we are running a variety of simulations using different combinations of protein and water models in order to evaluate the model-dependent differences in protein folding dynamics and resultant folded structures. Here we are focusing on 6 peptides/proteins: chignolin, trp-cage, bba, villin, ww domain, and homeodomain. Our goal is to determine which models produce the most accurate results.
SPECIFICATIONS
Uniform across projects:
x18 Nvidia GPU
8 runs
100 clones (1 us / clone)
250 gens (4 ns / gen)
10.8 deadline
2.0 timeout
0.75 k-factor
Unique for each project:
project protein natoms base-credit
p9606 chignolin 3575 130
p9607 chignolin 4720 190
p9608 trp-cage 5899 205
p9609 trp-cage 7770 310
p9610 bba 14104 690
p9611 bba 18636 1050
p9612 villin 9821 390
p9613 villin 12906 650
p9614 ww domain 15175 720
p9615 ww domain 20045 1180
p9616 homeodomain 11878 520
p9617 homeodomain 15530 830
Thank you  new x18 GPU projects {9606..9617}Beta team,
For this project series we are running a variety of simulations using different combinations of protein and water models in order to evaluate the model-dependent differences in protein folding dynamics and resultant folded structures. The goal is to figure out which models give the most accurate results.
SPECIFICATIONS
Uniform across projects:
x18 Nvidia GPU
8 runs
100 clones (1 us / clone)
250 gens (4 ns / gen)
10.8 deadline
2.0 timeout
0.75 k-factor
Unique for each project:
project protein natoms base-credit
p9606 chignolin 3575 120
p9607 chignolin 4720 200
p9608 trp-cage 5899 205
p9609 trp-cage 7770 310
p9610 bba 14104 690
p9611 bba 18636 1050
p9612 villin 9821 390
p9613 villin 12906 650
p9614 ww domain 15175 720
p9615 ww domain 20045 1180
p9616 homeodomain 11878 520
p9617 homeodomain 15530 830
|
|